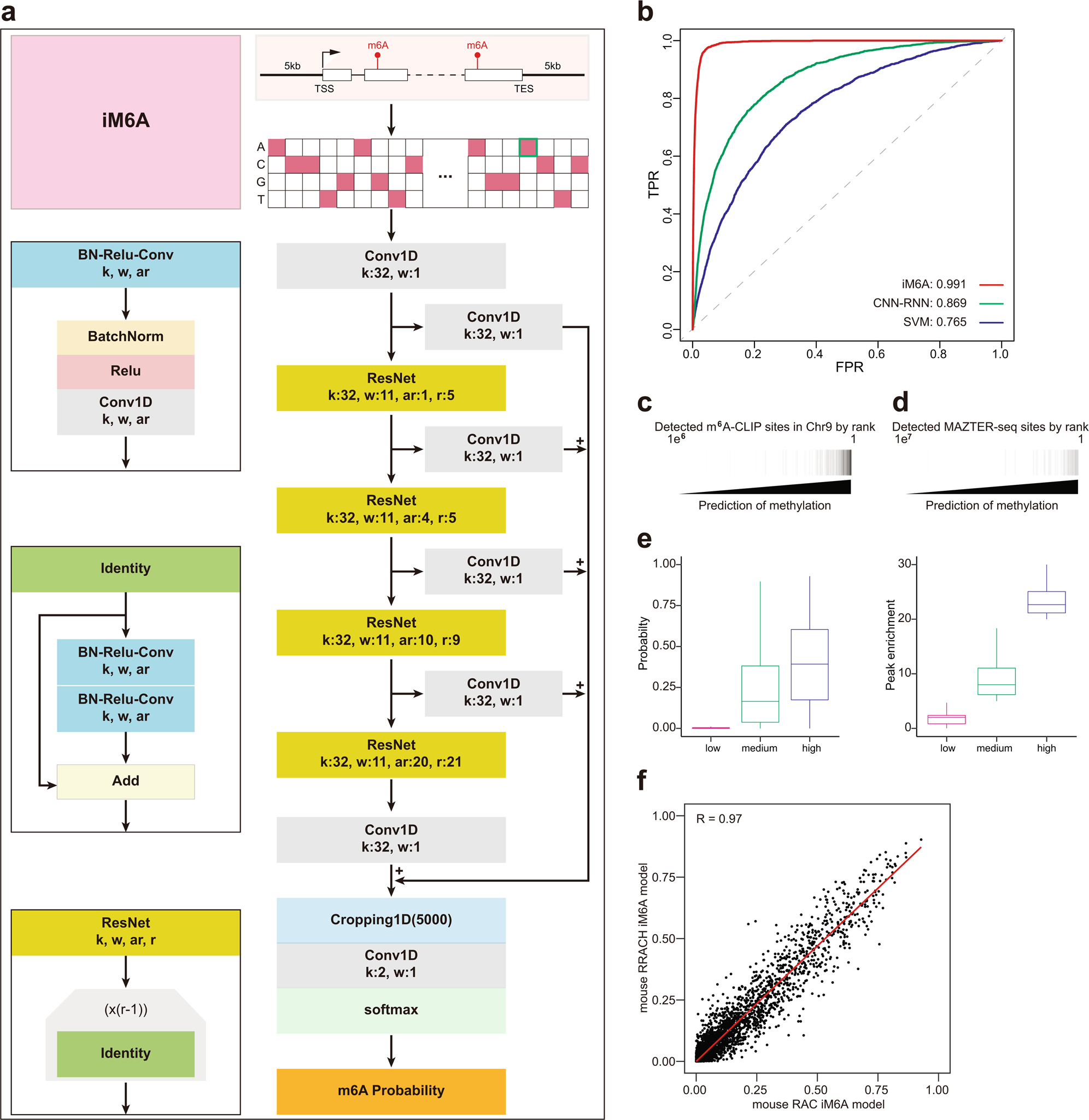
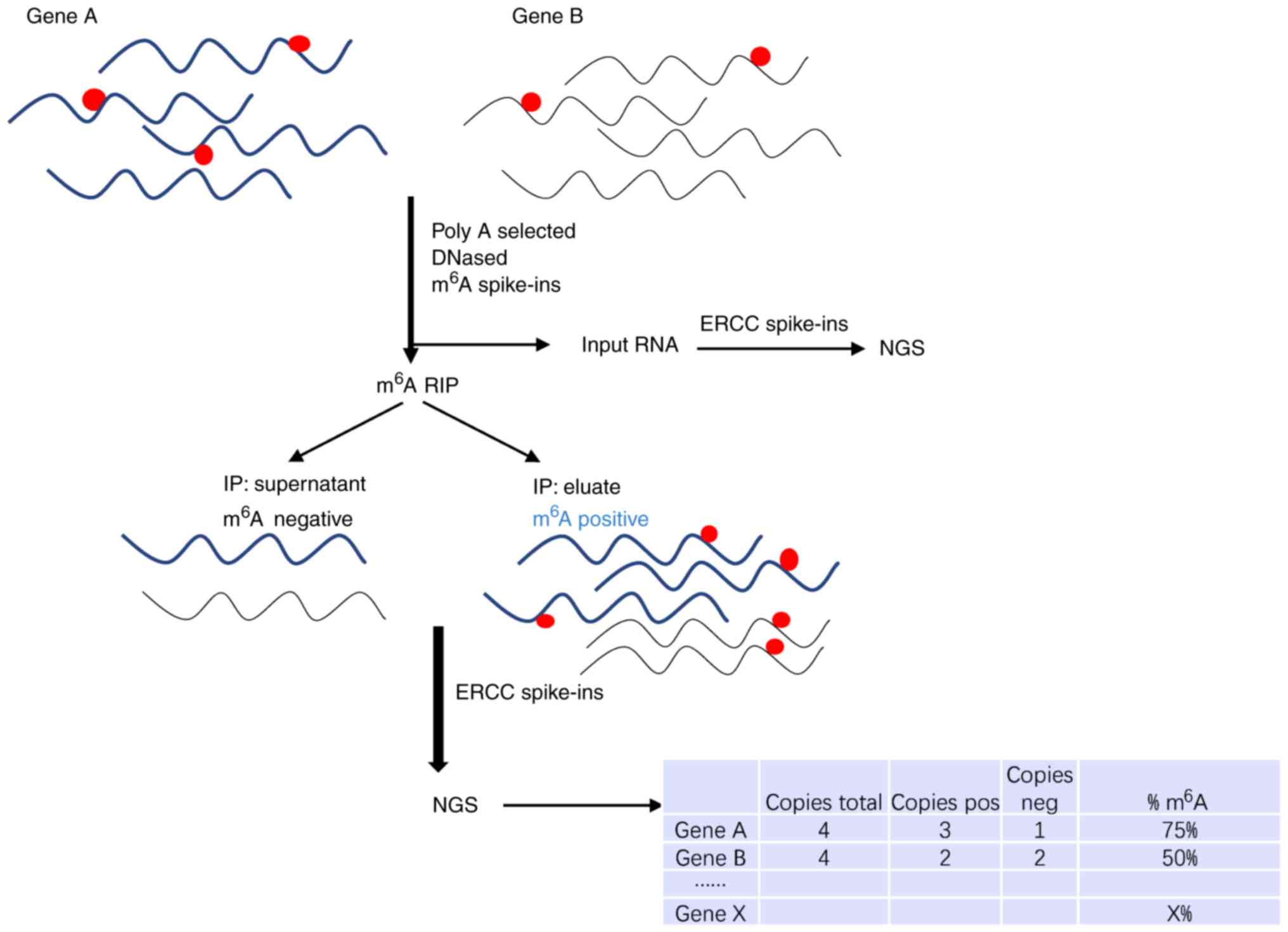
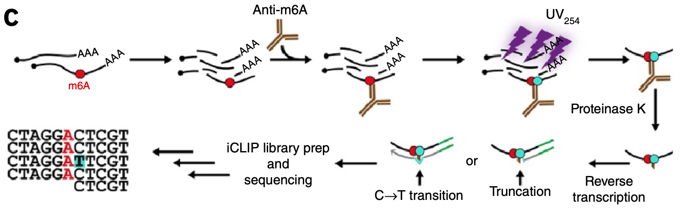
Frontiers | Emerging Perspectives of RNA N6-methyladenosine (m6A) Modification on Immunity and Autoimmune Diseases
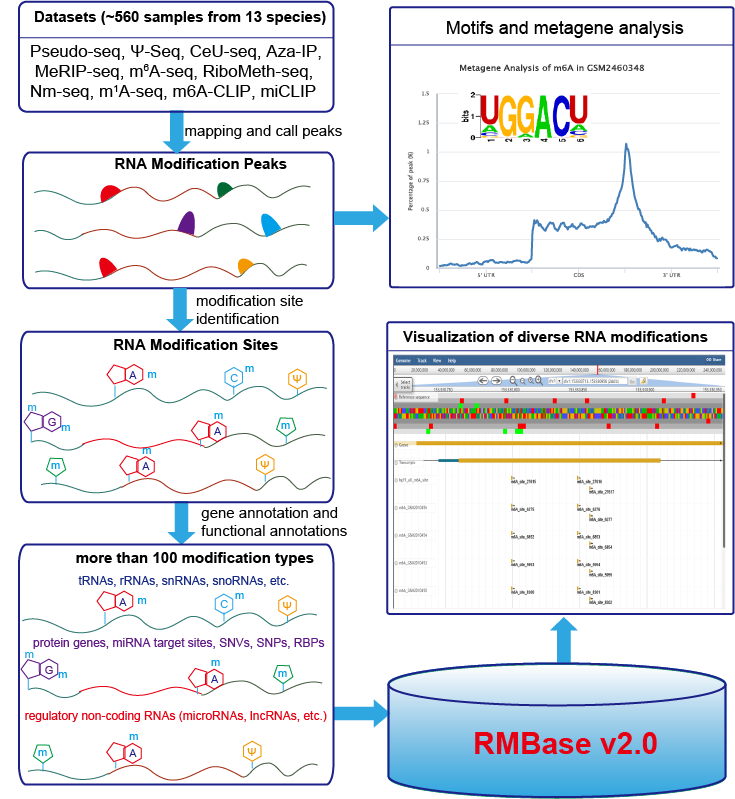
RNA Modification Database, RMBase v2.0: deciphering the map of RNA modifications from epitranscriptome sequencing data (Pseudo-seq, Ψ-seq, CeU-seq, Aza-IP, MeRIP-seq, m6A-seq, RiboMeth-seq, m1A-seq).

Antibody-free enzyme-assisted chemical approach for detection of N6-methyladenosine | Nature Chemical Biology

Single-nucleotide resolution achieved by m 6 A-CLIP/immunoprecipitation... | Download Scientific Diagram

The m6A reading protein YTHDF3 potentiates tumorigenicity of cancer stem-like cells in ocular melanoma through facilitating CTNNB1 translation | Oncogene
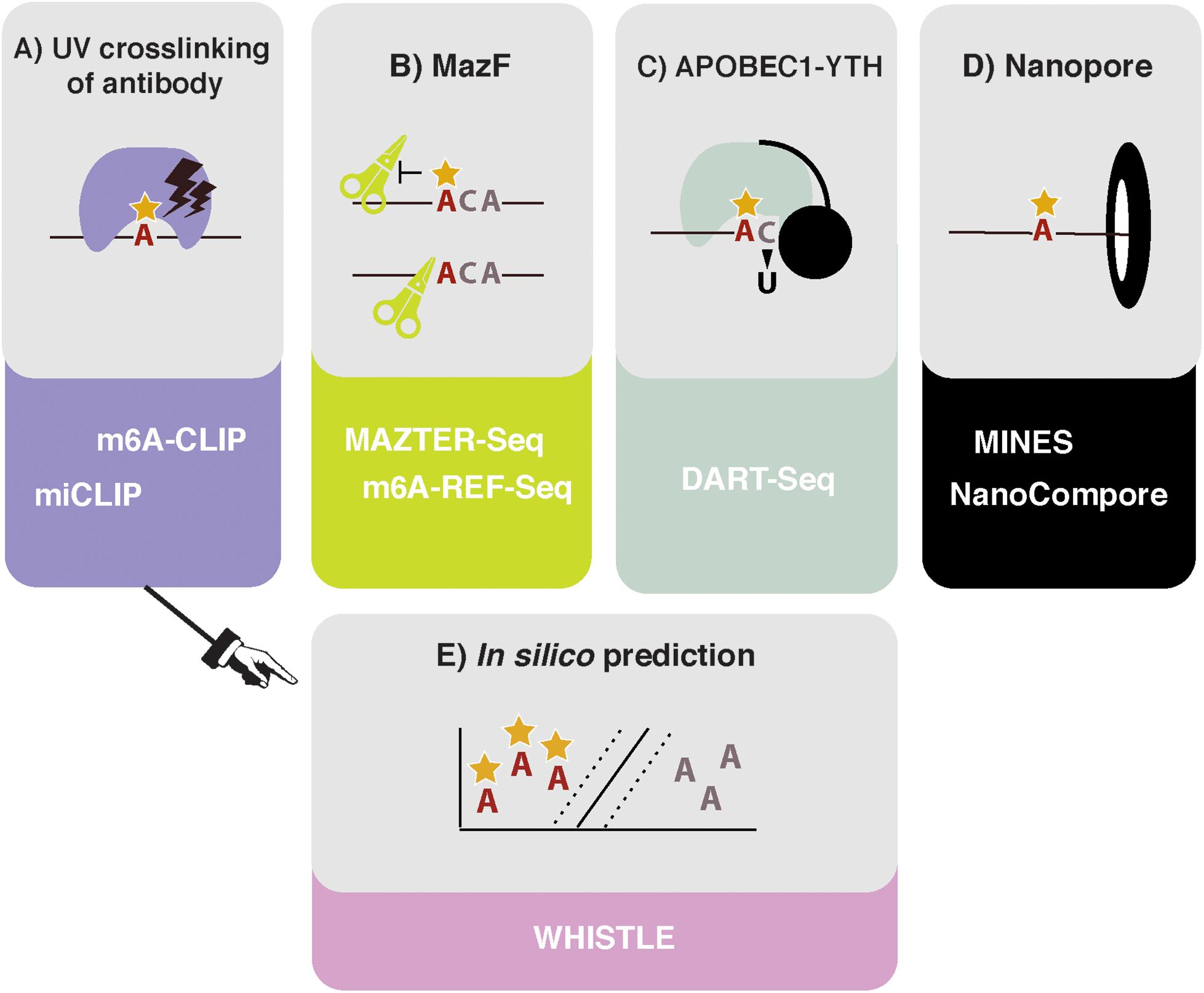
Frontiers | How Do You Identify m6 A Methylation in Transcriptomes at High Resolution? A Comparison of Recent Datasets

Down-Regulation of m6A mRNA Methylation Is Involved in Dopaminergic Neuronal Death | ACS Chemical Neuroscience

The detection and functions of RNA modification m6A based on m6A writers and erasers - ScienceDirect

The role of m6A modification in the biological functions and diseases | Signal Transduction and Targeted Therapy

Acute depletion of METTL3 identifies a role for N6-methyladenosine in alternative intron/exon inclusion in the nascent transcriptome | bioRxiv

m6A modification regulates lung fibroblast-to-myofibroblast transition through modulating KCNH6 mRNA translation: Molecular Therapy

Enhancer RNA m6A methylation facilitates transcriptional condensate formation and gene activation - ScienceDirect

Figure 2 from High-resolution N(6) -methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. | Semantic Scholar

Schematic diagram of miCLIP technology. m 6 A residues can be mapped by... | Download Scientific Diagram